ModelCube¶
Model Cube is a class to interact with a DAP model data cube for a galaxy.
Getting Started¶
ModelCube behaves in much the same way as a Cube. To initialize a ModelCube, you can specify either a mangaid, plateifu, or filename as input. Marvin will attempt to open the file locally from a file, a database, or remotely over the API.
from marvin.tools.modelcube import ModelCube
modelcube = ModelCube(mangaid='1-209232')
print(modelcube)
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='db', bintype='SPX', template='GAU-MILESHC')>
By default, it will grab the unbinned modelcube. You can specify a different binning with the bintype keyword.
modelcube = ModelCube(mangaid='1-209232', bintype='HYB10')
print(modelcube)
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='db', bintype='HYB10', template='GAU-MILESHC')>
You can quickly grab a spaxel/bin by slicing the ModelCube like an array.
# grab the bin of the central spaxel
bin_cen = modelcube[17, 17]
print(bin_cen)
<Marvin Bin (plateifu=8485-1901, x=17, y=17; x_cen=0, y_cen=0, n_spaxels=1)>
n_spaxels tells us there is only one spaxel in this bin. See Bin for more details on the Bin class. The binned`flux` in this bin is available as an attribute. It is represented as a Marvin Spectrum, which is a Quantity. To quickly plot the flux, use the plot method on the flux.
# look at the binned flux
bin_cen.flux
<Spectrum [ 0.54676276, 0.46566465, 0.4622981 ,..., 0. ,
0. , 0. ] 1e-17 erg / (Angstrom cm2 s spaxel)>
# plot the binned flux
bin_cen.flux.plot()
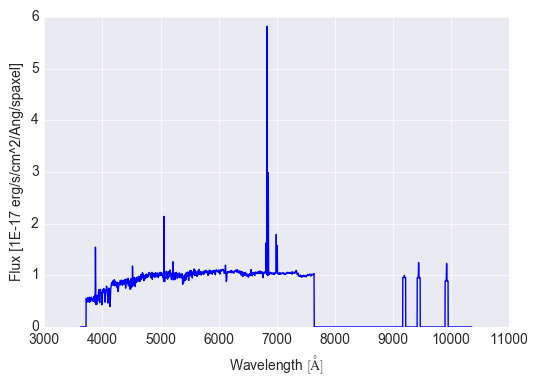
The full model fit is available as the full_fit attribute.
bin_cen.full_fit
<Spectrum [ 0., 0., 0.,..., 0., 0., 0.] 1e-17 erg / (cm2 s spaxel)>
# plot the model flux
bin_cen.full_fit.plot()
The ModelCube data quality and targeting flags are available as the quality_flag, target_flags, and pixmask attributes, respectively. These are represented as a Maskbit objects. A good quality ModelCube has an empty (0) bit list.
# check the quality and bits
modelcube.quality_flag
<Maskbit 'MANGA_DAPQUAL' []>
modelcube.quality_flag.bits
[]
# check the targeting flags
modelcube.target_flags
[<Maskbit 'MANGA_TARGET1' ['SECONDARY_v1_1_0', 'SECONDARY_COM2', 'SECONDARY_v1_2_0']>,
<Maskbit 'MANGA_TARGET2' []>,
<Maskbit 'MANGA_TARGET3' []>]
# retrieve the modelcube pixel mask
modelcube.pixmask
<Maskbit 'MANGA_DAPSPECMASK' shape=(4563, 34, 34)>
The DAPall information is accessible via the dapall attribute. It is a dictionary of the all the parameters from the DAPall file available for this target. Use dapall.keys() to see all of the available parameters.
# grab the star-formation rate within the IFU field-of-view
modelcube.dapall['sfr_tot']
0.132697
# and the mean surface brightness within 1 effective radius
modelcube.dapall['sb_1re']
0.738855
Using ModelCube¶
Initializing a ModelCube¶
A ModelCube can be initialized in several ways, by filename, in which case it will always be in local mode.
modelcube = ModelCube(filename='/Users/Brian/Work/Manga/analysis/v2_3_1/2.1.3/SPX-GAU-MILESHC/8485/1901/manga-8485-1901-LOGCUBE-SPX-GAU-MILESHC.fits.gz')
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='file', bintype='SPX', template='GAU-MILESHC')>
by plateifu or mangaid, in which case it attempts to find a local database, otherwise will open it in remote mode.
modelcube = ModelCube(plateifu='8485-1901', bintype='HYB10')
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='db', bintype='HYB10', template='GAU-MILESHC')>
modelcube = ModelCube(mangaid='1-209232', bintype='HYB10')
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='db', bintype='HYB10', template='GAU-MILESHC')>
However you can also initialize a ModelCube without the keyword argument and Marvin will attempt to figure out what input you mean.
modelcube = ModelCube('8485-1901', bintype='HYB10')
<Marvin ModelCube (plateifu='8485-1901', mode='local', data_origin='db', bintype='HYB10', template='GAU-MILESHC')>
Basic Attributes¶
Like ‘Cubes’, ModelCubes come with some basic attributes attached, e.g. the full header, the WCS info, the bintype and template, and the NSA and DAPall catalog parameters.
# access the header
modelcube.header
# access the wcs
modelcube.wcs
# the NSA catalog information
modelcube.nsa['z']
0.0407447
# the DAPall catalog info
modelcube.dapall['sfr_tot']
0.132697
ModelCube also has the DAP data quality, targeting, and pixel masks available as the quality_flag, target_flags, and pixmask attributes, respectively. These are represented as a Maskbit objects.
The DataModel¶
The DAP datamodel is attached to ModelCube as the datamodel attribute. The datamodel describes the contents of the MaNGA DAP ModelCube, for a given release. This is a subset of the full DAP datamodel only pertaining to ModelCubes.
# display the datamodel for DAP ModelCubes
modelcube.datamodel
[<Model 'binned_flux', release='2.1.3', unit=u'1e-17 erg / (cm2 s spaxel)'>,
<Model 'full_fit', release='2.1.3', unit=u'1e-17 erg / (cm2 s spaxel)'>,
<Model 'emline_fit', release='2.1.3', unit=u'1e-17 erg / (cm2 s spaxel)'>,
<Model 'emline_base_fit', release='2.1.3', unit=u'1e-17 erg / (cm2 s spaxel)'>]
Each Model describes its contents, units, and a description of what it is.
# look at the binned flux
modelcube.datamodel['binned_flux'].description
'Flux of the binned spectra'
# or the full_fit
modelcube.datamodel['full_fit'].description
'The best fitting model spectra (sum of the fitted continuum and emission-line models)'
Each Model also contains (and uses) the specific binid channel appropriate for that model. binned_flux and full_fit use the binned_spectra binids, while the emline models use the em_line_models binids.
modelcube.datamodel['binned_flux'].binid
<Property 'binid', channel='binned_spectra', release='2.1.3', unit=u''>
modelcube.datamodel['emline_fit'].binid
<Property 'binid', channel='em_line_models', release='2.1.3', unit=u''>
These are the available models used by DAP. Each Model is 3-d DataCube representation of the data within a DAP Cube. These models are available as attributes on your ModelCube object.
Models¶
All Models are DataCubes, which behave as Quantities.
# access the binned modelcube flux
modelcube.binned_flux
<DataCube [[[ 0., 0., 0.,..., 0., 0., 0.],
[ 0., 0., 0.,..., 0., 0., 0.],
[ 0., 0., 0.,..., 0., 0., 0.],
...,
[ 0., 0., 0.,..., 0., 0., 0.],
[ 0., 0., 0.,..., 0., 0., 0.],
[ 0., 0., 0.,..., 0., 0., 0.]]] 1e-17 erg / (cm2 s spaxel)>
The underlying numpy array data is always available as using the value attribute. They also may have available wavelength, ivar and mask attached.
# get the wavelength
modelcube.binned_flux.wavelength
<Quantity [ 3621.6 , 3622.43, 3623.26,..., 10349. , 10351.4 , 10353.8 ] Angstrom>
# get the ivar and mask as well
modelcube.binned_flux.ivar
modelcube.binned_flux.mask
If you slice the DataCube you get a Spectrum or another DataCube subset back.
# extract a single spectrum
modelcube.binned_flux[:,17,17]
<Spectrum [ 0.546763, 0.465665, 0.462298,..., 0. , 0. , 0. ] 1e-17 erg / (cm2 s spaxel)>
# extract a small cube around the center
subset_cen = modelcube.binned_flux[:,15:19,15:19]
print(subset_cen)
<DataCube [[[ 0.219631, 0.318331, 0.399484, 0.403951],
[ 0.288857, 0.419139, 0.517818, 0.552242],
[ 0.324734, 0.432396, 0.546763, 0.585823],
[ 0.310136, 0.395239, 0.486763, 0.48839 ]],
...,
[[ 0. , 0. , 0. , 0. ],
[ 0. , 0. , 0. , 0. ],
[ 0. , 0. , 0. , 0. ],
[ 0. , 0. , 0. , 0. ]]] 1e-17 erg / (cm2 s spaxel)>
Getting the Binids¶
For binned ModelCubes, you can retrieve a 2-d array of the binids using the get_binid method. For MPL-5, get_binid returns the binids from the BINID extension in the DAP files, while for MPL-6, by default, get_binid will return the binids for the binned_spectra channel of BINID.
# get the default binids
modelcube.get_binid()
array([[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
...,
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1]])
MPL-6 has new cubes using hybrid binning, **HYB10**, with alternate binning schemes. These are already built into the `Models`. `get_binid` can retrieve those with the `model` keyword.
::
# grab the binids for the emline_fit model
emline_binids = modelcube.get_binid(model=modelcube.datamodel['emline_fit'])
print(emline_binids)
array([[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
...,
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1],
[-1, -1, -1, ..., -1, -1, -1]])
Extracting Spaxels/Bins¶
If working with a unbinned ModelCube, slicing and getSpaxel will retrieve and return a Spaxel object, and behaves exactly the same as a Marvin Cube. For binned objects, slicing and extracting returns a Bin object instead. It behaves exactly the same as Spaxel except it now contains a list of spaxels belonging to that bin.
You can slice like an array
# slice a modelcube by i, j
bin_cen = modelcube[17, 17]
<Marvin Bin (plateifu=8485-1901, x=17, y=17; x_cen=0, y_cen=0, n_spaxels=1)>
# central bin id
bin_cen.binid
0.0
The central bin only contains on spaxel. Let’s go off-center.
# grab the bin for the array element 10, 10
newbin = modelcube[10,10]
print(newbin)
<Marvin Bin (plateifu=8485-1901, x=10, y=10; x_cen=-7, y_cen=-7, n_spaxels=20)>
# binid and bin SN
newbin.binid, newbin.bin_snr
(35.0, <AnalysisProperty 3.77872>)
This new bin has id 35, a signal-to-noise of ~4 and contains 20 spaxels. The spaxels attribute contains a list of all spaxels within this binid.
newbin.spaxels
[<Marvin Spaxel (x=9, y=10, loaded=False),
<Marvin Spaxel (x=9, y=11, loaded=False),
<Marvin Spaxel (x=10, y=8, loaded=False),
<Marvin Spaxel (x=10, y=9, loaded=False),
<Marvin Spaxel (x=10, y=10, loaded=False),
<Marvin Spaxel (x=10, y=11, loaded=False),
<Marvin Spaxel (x=10, y=12, loaded=False),
<Marvin Spaxel (x=11, y=9, loaded=False),
<Marvin Spaxel (x=11, y=10, loaded=False),
<Marvin Spaxel (x=11, y=11, loaded=False),
<Marvin Spaxel (x=11, y=12, loaded=False),
<Marvin Spaxel (x=12, y=8, loaded=False),
<Marvin Spaxel (x=12, y=9, loaded=False),
<Marvin Spaxel (x=12, y=10, loaded=False),
<Marvin Spaxel (x=12, y=11, loaded=False),
<Marvin Spaxel (x=12, y=12, loaded=False),
<Marvin Spaxel (x=13, y=9, loaded=False),
<Marvin Spaxel (x=13, y=10, loaded=False),
<Marvin Spaxel (x=13, y=11, loaded=False),
<Marvin Spaxel (x=13, y=12, loaded=False)]
Saving and Restoring¶
You can save a ModelCube locally as a Python pickle object, using the save method.
modelcube.save('mymodelcube.mpf')
as well as restore a ModelCube pickle object using the restore class method
from marvin.tools.modelcube import ModelCube
modelcube = ModelCube.restore('mymodelcube.mpf')
Reference/API¶
Class Inheritance Diagram
Class
marvin.tools.modelcube.ModelCube([input, …]) |
A class to interface with MaNGA DAP model cubes. |
Methods
marvin.tools.modelcube.ModelCube.get_binid([model]) |
Returns the 2D array for the binid map associated with model. |
marvin.tools.modelcube.ModelCube.get_unbinned() |
Returns a version of self corresponding to the unbinned ModelCube. |
marvin.tools.modelcube.ModelCube.getCube() |
Returns the associated Cube. |
marvin.tools.modelcube.ModelCube.getMaps() |
Returns the associated`~marvin.tools.maps.Maps`. |
marvin.tools.modelcube.ModelCube.getSpaxel([…]) |
Returns the Spaxel matching certain coordinates. |
marvin.tools.modelcube.ModelCube.download() |
Downloads the cube using sdss_access - Rsync |
marvin.tools.modelcube.ModelCube.save([…]) |
Pickles the object. |
marvin.tools.modelcube.ModelCube.restore(path) |
Restores a MarvinToolsClass object from a pickled file. |